Introduction
RT‑PCR (reverse transcription polymerase chain reaction) converts SARS‑CoV‑2 RNA into complementary DNA and amplifies specific viral gene targets, such as N, E, RdRp or S genes, allowing detection of a few copies of viral material. Because the assay amplifies genetic sequences, it achieves >95 % sensitivity and near‑100 % specificity when specimens—preferably nasopharyngeal swabs—are collected correctly, making it the gold‑standard diagnostic for active COVID‑19 infection. A positive result confirms current viral replication, while quantitative cycle‑threshold (Ct) values provide an indirect measure of viral load that correlates with infectivity and innate immune activation. By establishing the exact timing of infection, RT‑PCR guides the interpretation of serological tests, maps seroconversion kinetics, and supports vaccine‑efficacy studies, linking molecular detection directly to the host’s immune response and public‑health decision‑making.
1. RT‑PCR Is the Gold‑Standard Molecular Test

Mechanism of reverse transcription and amplification
RT‑PCR first converts SARS‑CoV‑2 RNA into complementary DNA (cDNA) using reverse transcriptase.
The cDNA then serves as a template for polymerase chain reaction, where specific primers (targeting N, E, RdRp, or S genes) and fluorescent probes amplify the viral genome exponentially.
Real‑time detection of fluorescence after each cycle yields a cycle‑threshold (Ct) value, allowing even a single viral copy to be identified.
High sensitivity and specificity
Because of this amplification, RT‑PCR detects as few as 10‑100 viral RNA copies per reaction, giving analytical sensitivity >95 % when specimens are collected correctly.
Specific primers and probes ensure specificity approaching 100 %, minimizing false‑positives.
Lower Ct values reflect higher viral loads, correlating with infectivity and disease severity, which is essential for clinical and public‑health decisions.
Accreditation standards (NABL, ICMR)
Laboratories such as Agam Diagnostics follow NABL and ICMR accreditation adhering to stringent quality‑control protocols, internal controls, and external proficiency testing.
These standards guarantee reproducible performance, rapid turnaround (≤24 hours) and reliable results for diagnosing active COVID‑19 infection and supporting downstream immunological studies.
2. Sample Collection: The Key to Accurate Results

Nasopharyngeal (NP) swabbing is the gold‑standard for COVID‑19 RT‑PCR because the posterior nasopharynx harbors the highest viral load during the first 5‑7 days of infection. A correctly performed NP swab reaches this region, is rotated for at least 10 seconds, and is placed immediately in viral transport medium; this maximises the amount of RNA available for amplification and drives the assay’s high sensitivity (>95%). Sub‑optimal collection—shallow nasal or oropharyngeal swabs, insufficient rotation, or delayed transport—lowers viral RNA yield, increasing the risk of false‑negative results, especially when testing is done very early (≤3 days post‑exposure) or after viral clearance. Trained phlebotomists at Agam Diagnostics follow strict protocols and temperature‑controlled logistics to preserve RNA integrity, ensuring reliable RT‑PCR outcomes and supporting accurate immunological assessments.
3. Timing of Testing and False‑Negative Risks

RT‑PCR is most reliable when viral RNA concentrations peak, which occurs around day 5‑6 after symptom onset or 5‑7 days after exposure. During this window the assay’s limit of detection (10‑100 copies) is comfortably exceeded, yielding high sensitivity and specificity. Testing too early—within the first 0‑3 days of exposure—often captures viral loads below this threshold, leading to false‑negative results. Likewise, testing after viral clearance can miss infection entirely. Because early testing can miss infection, clinical guidelines recommend repeat testing after 24‑48 hours when an initial negative result is obtained from a symptomatic individual or a high‑risk contact. This repeat approach improves diagnostic confidence and ensures timely isolation, contact tracing, and initiation of therapy, especially for vulnerable populations such as the immunocompromised.
4. Cycle‑Threshold (Ct) Values: Viral Load and Immune Response

RT‑PCR cycle‑threshold (Ct) values are the point at which fluorescence from amplified viral DNA exceeds background. Because each cycle roughly doubles the target, a lower Ct indicates that fewer cycles were needed to detect signal, reflecting a higher concentration of SARS‑CoV‑2 RNA in the specimen. In practice, Ct ≤ 20‑25 corresponds to a high viral load, while Ct > 30‑35 denotes low RNA levels. This quantitative surrogate correlates with several immunological parameters. Patients with low Ct values often exhibit stronger innate immune activation, particularly type I interferon responses, and are more likely to be infectious. Moreover, high viral loads accelerate the kinetics of adaptive immunity: IgM antibodies appear sooner and IgG seroconversion follows earlier, providing a temporal map of the host’s humoral response. Conversely, rising Ct values over time signal decreasing viral burden, coinciding with the resolution of cytokine spikes (e.g., IL‑6) and a shift toward recovery. Although Ct is not uniformly calibrated across platforms, it remains a valuable indicator of disease severity, transmissibility, and the timing of immune events.
5. RT‑PCR in Serology and Vaccine Efficacy Studies

RT‑PCR is the gold‑standard method for confirming active SARS‑CoV‑2 infection before any serological testing. Because IgM and IgG antibodies appear only days after viral replication begins, a positive RT‑PCR result provides the precise infection onset needed to map seroconversion kinetics. This temporal anchor lets researchers link cycle‑threshold (Ct) values and viral load peaks with the timing of antibody emergence, distinguishing vaccine‑induced immunity from natural infection. In vaccine efficacy studies, RT‑PCR also identifies breakthrough infections by verifying that a new viral RNA signal reflects a genuine infection rather than lingering RNA fragments, allowing accurate calculation of vaccine protection against emerging variants. Combining RT‑PCR data with immunology panels (e.g., cytokine profiling, neutralizing antibody titers) offers a comprehensive picture of both active infection status and the host’s immune response, guiding personalized treatment and public‑health strategies.
6. Special Considerations for Immunocompromised Patients

immunocompromised individuals often experience delayed viral clearance and can shed SARS‑CoV‑2 for weeks, making early detection critical. RT‑PCR’s high analytical sensitivity (detecting as few as 10–100 copies of viral RNA) enables identification of low‑level infection even before symptoms appear, allowing prompt isolation and initiation of targeted antivirals such as remdesivir or monoclonal antibodies. Serial RT‑PCR monitoring provides quantitative Ct values that reflect viral load kinetics; falling viral loads guide the duration of antiviral therapy and help decide when to add immunomodulatory agents like corticosteroids or IL‑6 blockers. By confirming active infection and tracking viral persistence, RT‑PCR reduces the risk of severe disease progression and informs personalized treatment strategies for this vulnerable population.
7. Integrated Immunology Panels at Agam Diagnostics

Agam Diagnostics combines its gold‑standard RT‑PCR COVID detectionplatform with a full suite immun immunology testing, delivering a comprehensive picture of both viral load and host response. The laboratory’s fully automated, NABL‑ and ICMR‑accredited RT‑PCR system provides results within 24 hours, and the same specimen can be used for quantitative cytokine profiling (e.g., IL‑6, CRP), lymphocyte subset analysis, and antibody titers. By linking Ct‑derived viral load values with these immunological biomarkers, clinicians can rapidly assess disease severity, predict progression, and tailor interventions such as antiviral therapy, immunomodulatory drugs, or booster vaccination. This integrated approach shortens the diagnostic timeline, reduces the need for multiple referrals, and supports timely, personalized clinical decision‑making.
Conclusion
In summary, nine essential facts about RT‑PCR for COVID‑19 emerge from the evidence: (1) it is the gold‑standard molecular test, detecting SARS‑CoV‑2 RNA with >95% sensitivity and near‑perfect specificity when specimens are properly collected. (2) Nasopharyngeal swabs reaching the posterior nasopharynx yield the highest viral load and minimize false‑negatives. (3) Viral load peaks 5‑6 days after symptom onset, so testing within this window maximizes detection. (4) Cycle‑threshold (Ct) values provide an indirect viral‑load measure, informing infectivity and disease severity. (5) Early RT‑PCR positivity precedes antibody seroconversion, guiding timing of serology. (6) False‑negatives arise from early testing, poor sampling, or low viral levels, warranting repeat tests. (7) RT‑PCR results drive rapid isolation, contact tracing, and therapeutic decisions. (8) Integration with immunology panels (cytokines, antibodies) yields a comprehensive view of host response. (9) High‑throughput, accredited laboratories (e.g., Agam Diagnostics) deliver ≤24‑hour turnaround, supporting both individual care and population‑level surveillance, underscoring RT‑PCR’s pivotal role in immunology research and public‑health strategy.
Why Microbiology Matters in Emerging Disease Surveillance
Surveillance is the continuous, systematic collection, analysis and interpretation of health data to guide public‑health planning, implementation and evaluation. Clinical microbiology laboratories have been the first point of detection for infectious threats since the advent of culture and microscopy, evolving from simple Gram stains to sophisticated molecular platforms that can sequence entire pathogen genomes in hours. Emerging threats—novel viruses such as SARS‑CoV‑2, influenza A variants, bioterrorism agents, and zoonotic spillovers like Nipah or Streptococcus suis—underscore the need for rapid, accurate laboratory data. Modern labs, exemplified by fully automated facilities such as Agam Diagnostics in Madurai, provide same‑day PCR, whole‑genome sequencing, and automated alert systems that feed into regional, national and international surveillance networks, enabling early intervention and containment.
Surveillance Foundations and the Role of Microbiology Laboratories

Surveillance is the continuous, systematic collection, analysis, and interpretation of health data that underpins public‑health planning, implementation, and evaluation.
Clinical microbiology laboratories serve as the first detection point for emerging threats and feed data into three surveillance models. Passive surveillance relies on routine, often paper‑based, analysis of all microbiological results, generating alerts after the fact. Passive surveillance relies on routine analysis of all microbiological data and is often paper‑based. Active surveillance applies predefined thresholds and engages trained microbiologists and epidemiologists to seek out abnormal patterns, enabling faster outbreak detection. Active surveillance applies predefined thresholds and engages trained microbiologists and epidemiologists. Virtual surveillance combines active data collection with advanced computing, mathematical models, and real‑time analytics to flag atypical pathogen trends instantly. Virtual surveillance combines active surveillance with advanced computing technologies and mathematical models to provide real‑time detection of abnormal pathogen patterns. Automated laboratory information systems in microbiology labs generate electronic alerts, stream‑ data to regional, national, and international networks such as WHO GLASS, WHONET, and national Integrated Disease Surveillance Programs. Automated data management systems in microbiology labs can generate alerts for unusual results, detect trends, and integrate data into regional, national, and international surveillance networks. By integrating specimen management, rapid molecular diagnostics, and robust data pipelines, microbiology laboratories enhance temporal and spatial disease mapping, support early intervention, and improve coordinated public‑health responses worldwide.
Emerging Pathogens and the Need for Rapid Detection

Rapid detection of emerging pathogens such as SARS, novel influenza A variants, bioterrorism agents, and zoonotic spillovers is now a public‑health imperative. Delayed recognition allows unchecked transmission, inflating outbreak size and mortality; the 2002‑2003 SARS epidemic and the 2001 anthrax mail attacks both demonstrated how weeks of diagnostic lag amplified case numbers and panic. Recent laboratory data illustrate the stakes: SARS‑CoV‑2 has been isolated from white‑tailed deer in Ohio, indicating wildlife reservoirs that can reseed human infections if not identified early. Likewise, the emergence of OXA‑48‑producing Escherichia coli in food‑premises highlights how antibiotic‑resistant bacteria can spread silently through the food chain, evading routine culture‑based surveillance. Real‑time PCR, whole‑genome sequencing, and automated data‑alert systems—implemented in accredited facilities such as Agam Diagnostics—provide same‑day pathogen identification, resistance‑gene profiling, and rapid reporting to national and international networks. These technologies shrink turnaround times from days to hours, enabling timely isolation, targeted therapy, and coordinated public‑health responses that curb the magnitude of emerging infectious disease threats.
Molecular Diagnostics: PCR, Real‑time PCR, and Next‑Generation Sequencing
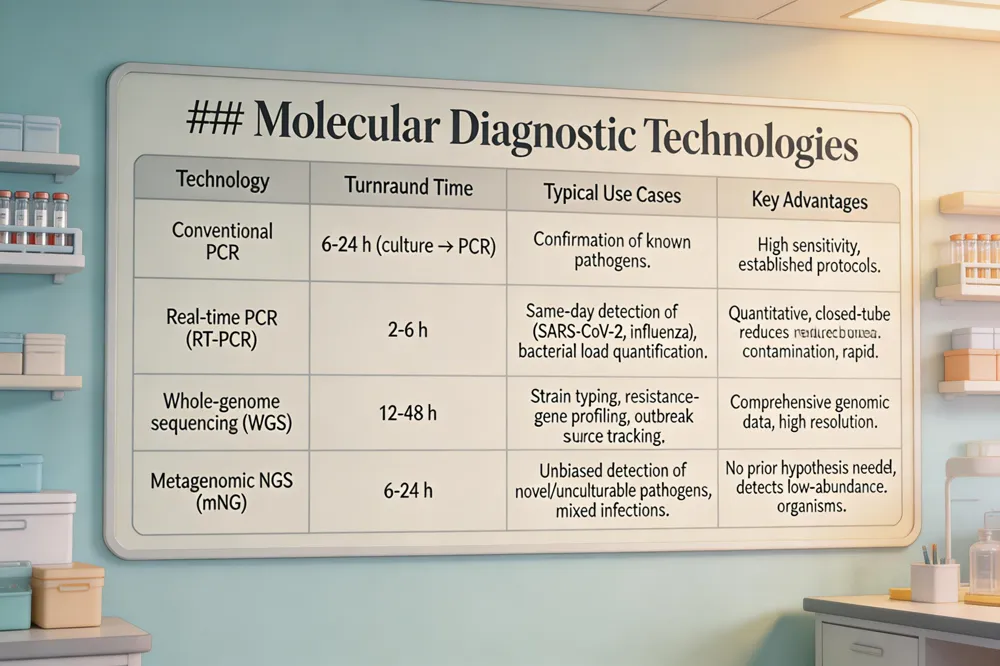

Real‑time PCR (RT‑PCR) amplifies nucleic acids while simultaneously measuring fluorescence, delivering quantitative results in 2‑6 hours. Its closed‑tube format minimizes contamination and, combined with automated extraction, shortens turnaround from days (culture) to hours, enabling same‑day detection of emerging viruses such as SARS‑CoV‑2, influenza A variants, and zoonotic agents. Whole‑genome sequencing (WGS) and metagenomic next‑generation sequencing (mNGS) go further by reading the entire genetic material of a specimen without prior culturing. These platforms identify novel or unculturable pathogens, map mutations that affect virulence, and reveal transmission pathways in real time. In antimicrobial‑resistance surveillance, WGS pinpoints resistance genes (e.g., OXA‑48, mcr‑1) and plasmid backgrounds, while mNGS can detect low‑abundance resistant strains in mixed infections. Together, RT‑PCR provides rapid, targeted alerts for known threats, whereas WGS/mNGS supplies comprehensive genomic data for strain tracking, outbreak source attribution, and informed public‑health interventions.
Automation and Core Laboratory Models for Speed and Cost‑Effectiveness

Horizontal core laboratory structures bring microbiology, immunology and biochemistry under a single, unified workflow, eliminating duplicated steps and streamlining staffing. By consolidating these disciplines, laboratories can share high‑throughput analyzers, automated incubators and common data‑management platforms, which reduces capital outlay and improves turnaround time for all test types. Robotic specimen handling—cover bar‑code‑driven sorting, automated aliquoting and sealed biosafety cabinets—paired with an integrated Laboratory Information System (LIS) enables real‑time data capture, trend detection and automatic alert generation for unusual results. These alerts feed directly into regional and national surveillance networks, shortening reporting from days to hours. In high‑volume settings such as Agam Diagnostics in Madurai, the model yields measurable cost savings through lower consumable waste, fewer repeat tests and optimized personnel allocation, while delivering same‑day pathogen identification and antimicrobial‑susceptibility results essential for outbreak control.
Specimen Management and Biosafety Best Practices

Effective specimen management begins with proper collection: trained staff must use sterile containers, follow pathogen‑specific protocols, and label each sample with patient identifiers, date, time, and test request. Immediate, temperature‑controlled transport—refrigerated for most bacterial and viral specimens, frozen for RNA viruses—preserves nucleic‑acid integrity, while prompt storage at the lab prevents degradation. In the microbiology laboratory, Class II biosafety cabinets provide a protected environment for handling potentially infectious material; personnel must wear appropriate personal protective equipment (gloves, lab coats, eye protection, and N95 respirators when aerosol‑generating procedures are performed. Waste is decontaminated by autoclaving or chemical disinfection before disposal. High‑quality specimens are essential: contaminated or poorly preserved samples yield false‑negative results, undermine antimicrobial‑resistance surveillance, and delay outbreak detection. By standardising collection, transport, and biosafety measures, laboratories such as Agam Diagnostics ensure accurate diagnostics and reliable data for public‑health surveillance.
Data Integration, Automated Alerts, and Virtual Surveillance
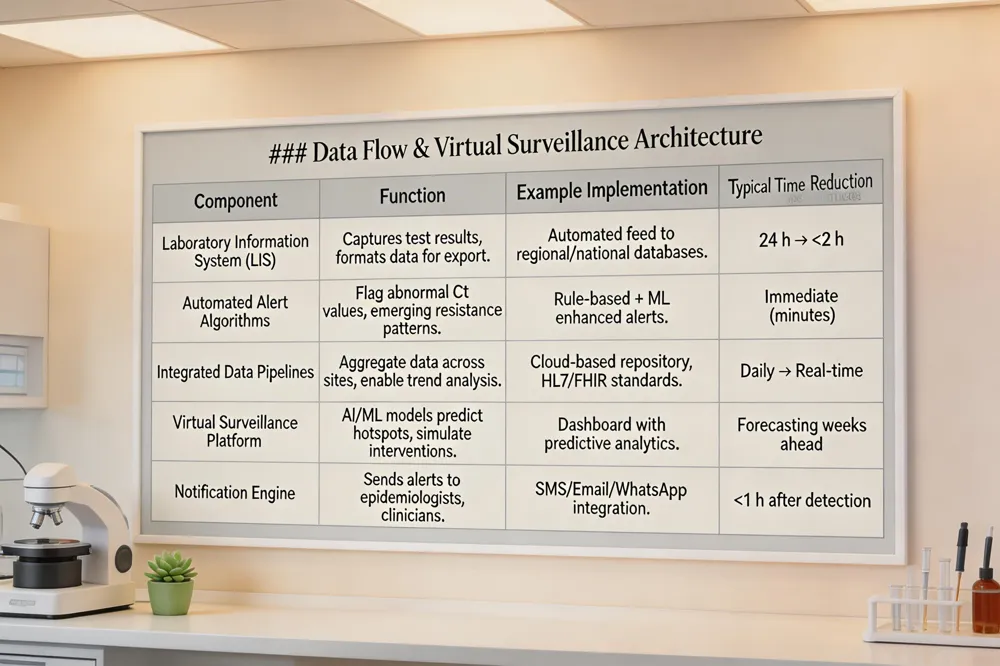

Modern clinical microbiology laboratories, such as Agam Diagnostics in Madurai, now employ robust Laboratory Information Systems (LIS) that automatically feed test results into regional, national, and supranational surveillance networks (e.g., WHO GLASS, WHONET). These electronic links enable near‑real‑time aggregation of pathogen data, allowing automated alert algorithms to flag unusual results—such as unexpected viral PCR Ct values or emergence of carbapenem‑resistant bacterial isolates—within hours of receipt. Integrated data pipelines support trend analysis across time and space, facilitating early outbreak detection and targeted public‑health interventions. Building on this foundation, virtual surveillance platforms incorporate mathematical modeling and artificial‑intelligence tools to predict abnormal pathogen patterns, assess transmission dynamics, and simulate the impact of control measures. AI‑driven dashboards can prioritize alerts, suggest laboratory actions, and automatically notify epidemiologists and clinicians, thereby shortening reporting time from a median of five days to as little as one day. Together, LIS connectivity, automated alerts, and AI‑backed virtual surveillance create a rapid, data‑centric defense against emerging infectious disease threats.
Agam Diagnostics: Infrastructure, Accreditation, and Service Reach

Agam Diagnostics is a fully automated pathology laboratory situated in Madurai, Tamil Nadu, India. It operates a high‑throughput, integrated platform that combines microbiology, molecular biology, haematology, clinical biochemistry, immunology and medical genetics under one roof, enabling same‑day or next‑day reporting for most assays. The laboratory holds NABL accreditation (ISO 15189:2022) and is also accredited by the Indian Council of Medical Research (ICMR), ensuring compliance with stringent quality‑assurance programmes, regular participation in external quality‑assessment schemes, and a documented 99.9 % accuracy rate for test results. To maximise accessibility, Agam Diagnostics offers free home‑sample collection across Tamil Nadu and a network of partner laboratories in seven additional locations, ensuring timely specimen transport and rapid turnaround even for patients in remote areas. This combination of automation, accreditation, and outreach positions the lab as a critical node for early detection of emerging infectious diseases and for feeding reliable data into national surveillance systems.
Impact on Public Health: Early Intervention, Outbreak Control, and Antimicrobial Resistance Monitoring

Temporal and spatial analysis of disease patterns becomes possible when clinical microbiology laboratories feed high‑quality, time‑stamped data into national surveillance networks. Automated laboratory information systems generate alerts for unusual pathogen trends, allowing health authorities to map hotspots and predict spread across districts. Rapid antimicrobial susceptibility testing (AST) shortens the interval from specimen receipt to therapeutic decision: hours rather than days, enabling clinicians to prescribe targeted agents and curb the use of broad‑spectrum antibiotics that drive resistance. In India, laboratories such as Agam Diagnostics, which are NABL and ICMR accredited, integrate their AST results with the Integrated Disease Surveillance Programme (IDSP) and the WHO Global Antimicrobial Resistance Surveillance System (GLASS). This linkage provides real‑time resistance profiles at the regional and national levels, supporting evidence‑based treatment guidelines, informing infection‑control policies, and enhancing early outbreak response. The combined effect is a more agile public‑health system that can intervene promptly, limit transmission, and monitor antimicrobial resistance trends effectively.

Unbiased pathogen detection through shotgun metagenomics will soon become routine in microbiology labs such as Agam Diagnostics, allowing clinicians to identify known, novel, or unculturable agents directly from clinical specimens without prior hypotheses. Coupled with rapid next‑generation sequencing platforms, this approach can pinpoint viral variants, antimicrobial‑resistance genes, and zoonotic spill‑over events within hours, feeding real‑time data into regional and national surveillance networks. Machine‑learning algorithms will ingest these high‑dimensional datasets, together with electronic laboratory reporting and epidemiological indicators, to predict outbreak hotspots, forecast resistance trends, and generate automated alerts for unusual pathogen signatures. The One Health model further expands the data stream by integrating human, animal, and environmental microbiology results—such as wildlife sampling for SARS‑CoV‑2 variants or livestock testing for emerging bacterial strains—into a unified analytical framework. Together, metagenomics, AI, and One Health surveillance promise faster, more precise public‑health responses to emerging infectious diseases.
Putting It All Together – The Critical Role of Microbiology in a Changing Infectious‑Disease Landscape
Clinical microbiology laboratories provide the first line of detection for emerging threats, turning specimen collection, culture, microscopy, antigen assays, and molecular tests into actionable data. Rapid diagnostics—real‑time PCR, next‑generation sequencing, MALDI‑TOF—identify pathogens within hours, allowing clinicians to start appropriate therapy and public‑health agencies to trigger outbreak alerts. Agam Diagnostics exemplifies this modern model: a NABL‑ and ICMR‑accredited, fully automated facility in Madurai that offers bacterial culture, viral PCR panels, antimicrobial‑susceptibility testing, and free home sample collection. Its integrated laboratory information system generates electronic alerts and feeds results into regional surveillance networks, shortening reporting times from days to a single day. To sustain these gains, continued investment is needed in cutting‑edge platforms, regular staff training, and robust data‑sharing frameworks that link microbiology labs with epidemiologists and national surveillance programs. Such support will keep the laboratory’s early‑warning capability sharp as pathogens evolve and spread.
Introduction
Telemedicine, once limited to telephone triage, has rapidly evolved into a comprehensive digital health ecosystem that supports real‑time video consultations, remote monitoring, and electronic ordering of laboratory tests. The COVID‑19 pandemic acted as a catalyst, accelerating the deployment of home‑testing kits, expanding broadband access, and prompting regulatory bodies to clarify reimbursement and privacy standards. Key drivers of home‑testing adoption include the need to reduce travel for patients in rural India, the convenience of free phlebotomy services, and the ability to obtain rapid, laboratory‑grade results without visiting a clinic. Automated pathology laboratories such as Agam Diagnostics in Madurai play a pivotal role: their NABL‑ and ICMR‑accredited, high‑throughput platforms deliver results within 24‑48 hours, while integrated electronic health information systems enable seamless order transmission and secure result delivery. Together, these advances close the gap between virtual care and definitive diagnostics, improving access, lowering costs, and supporting timely clinical decisions.
Remote Diagnosis: Technologies and Clinical Impact

Remote diagnosis in telemedicine relies on a suite of digital tools that bring laboratory‑grade testing to the patient’s home. Digital imaging and secure cloud platforms enable clinicians to view high‑resolution scans of blood smears, radiographs, or histopathology slides uploaded from automated labs such as Agam Diagnostics; encryption and HIPAA‑level safeguards protect patient data while allowing instant specialist review. Wearable biosensors and real‑time data streaming provide continuous vital‑sign feeds—glucose, blood pressure, SpO₂, ECG—directly to telehealth dashboards, supporting early detection of deterioration and informing when a home‑collection kit is needed. AI‑driven decision‑support and analytics process these streams and laboratory results, flagging abnormal patterns, calculating risk scores, and suggesting evidence‑based test panels, thereby boosting diagnostic accuracy and reducing clinician workload. Telemedicine operates in two interaction modes: live interactive (real‑time video, synchronous data exchange) permits immediate clinical assessment and on‑the‑spot ordering of home‑collection kits, while store‑and‑forward (asynchronous transmission of images, lab orders, and sensor logs) allows specialists to review cases at their convenience, expanding reach to rural settings where bandwidth may be limited. Together, these technologies shorten turnaround times—often delivering results within 24‑48 hours—improve access for underserved populations, and lay the groundwork for scalable, equitable remote care.
Agam Diagnostics: Bridging Rural Gaps with Home Collection

Agam Diagnostics expands tele‑medicine reach in India by offering free at phlebotomy that brings a qualified phlebotomist to the patient’s doorstep, collects blood, urine or swab specimens, and transports them in temperature‑controlled containers to its fully automated pathology laboratory in Madurai. The laboratory’s high‑throughput, robotic workflow processes hundreds of samples simultaneously, minimizing manual handling and reducing the risk of human error. Because the facility is Accredited by the National Accreditation Board for Testing and Calibration Laboratories (NABL) and follows Indian Council of Medical Research (ICMR) guidelines, every test meets internationally recognized quality standards. This integrated model delivers routine panels—such as complete blood counts, metabolic profiles and lipid panels—within 24‑48 hours, enabling clinicians to review results in real time during virtual consultations and make timely treatment decisions for patients in remote or underserved communities.

Remote diagnostic models are evaluated with the same rigor as in‑person testing. Core performance measures include accuracy, sensitivity, specificity, area under the ROC curve (AUC) and F1‑score, which together quantify how well a telemedicine‑driven test identifies disease, avoids false positives, and balances precision with recall. In the telehealth workflow, turnaround time has emerged as a critical clinical KPI; rapid delivery of results—often within 24‑48 hours at fully automated labs such as Agam Diagnostics—enables timely therapeutic decisions and reduces patient anxiety. Automation underpins this speed: high‑throughput analyzers, robotic sample handling, and AI‑assisted quality control lower manual handling, reduce transcription errors, and improve reproducibility across large test volumes. Seamless integration of laboratory information systems (LIS) with telehealth electronic medical records (EMR) completes the quality loop. Orders are transmitted electronically, specimen metadata are captured at the point of home collection, and results flow back through encrypted channels, ensuring data integrity, traceability, and compliance with NABL and ICMR standards. Together, these metrics and technological safeguards guarantee that remote testing delivers reliable, rapid, and equitable care.
Challenges and Future Directions: Data Privacy, Integration, Equity

Remote diagnosis in telemedicine promises rapid, convenient care, but several challenges must be addressed before it can achieve its full potential.
Data security and compliance – Indian regulations such as the IT Act and upcoming Personal Data Protection Bill require encrypted transmission and storage of patient information. Platforms that link telehealth visits with labs like Agam Diagnostics must use HTTPS, HIPAA‑style safeguards, and cloud‑based data rooms that meet both national (NABL, ICMR) and international standards to protect privacy and maintain patient trust.
Heterogeneous data source integration – A modern telehealth workflow blends vitals from wearable sensors (e.g., Bluetooth‑connected glucometers, blood‑pressure cuffs), high‑resolution images (digital pathology slides, chest X‑rays) and laboratory results from automated centers. Seamless integration demands interoperable standards (FHIR, HL7) and robust laboratory information systems that can ingest and reconcile disparate data streams without loss of fidelity.
Real‑time analytics and AI explainability – AI‑driven decision‑support tools are increasingly used to flag abnormal lab values or to pre‑screen digital pathology images. While such algorithms can raise diagnostic accuracy (up to 91% in AI‑enhanced telemedicine), clinicians need transparent, explainable outputs to avoid over‑reliance and to comply with regulatory expectations for auditability.
Equitable access across rural and urban populations – Over 65% of India's population lives in rural areas where broadband connectivity, digital literacy, and device availability are limited. Initiatives such as free home phlebotomy (Agam Diagnostics), low‑cost mHealth apps, and government‑backed National Digital Health Mission aim to bridge this gap, but sustained investment in infrastructure, training, and reimbursement models is essential to prevent a new digital divide.
Economic and Public Health Impact: Cost Savings and Access Expansion

Remote diagnosis and home‑collection services such as those offered by Agam Diagnostics dramatically cut patient‑borne costs. By eliminating the need for travel to a physical laboratory, patients save on transportation expenses, lost wages, and ancillary costs such as childcare or accommodation, especially in rural India where 70% of the population lives far from urban health centres. This financial relief is amplified when telemedicine platforms shift routine follow‑up diagnostics away from hospitals, reducing overcrowding and freeing inpatient capacity for acute cases. Chronic disease management benefits equally; continuous monitoring devices (glucometers, blood‑pressure cuffs, wearable sensors) feed real‑time data to clinicians who can order timely laboratory tests via telehealth, enabling early detection of deterioration and preventing costly hospital admissions. The broader market reflects these efficiencies: a Systematic review (2016‑2023) notes that telemedicine reduces overall healthcare spending, while industry forecasts project the global telehealth market to reach approximately USD 36.5 billion by 2032 with a 17.3% CAGR (2023‑2032). Together, cost savings, reduced crowding, and improved chronic‑care pathways illustrate how remote diagnostic ecosystems expand access while delivering measurable economic benefits.
Conclusion
The convergence of telemedicine platforms with Agam Diagnostics creates a seamless, end‑to‑end remote diagnostic pathway. Clinicians can order tests through virtual visits, patients receive free home phlebotomy, and the fully automated, NABL‑ and ICMR‑accredited laboratory delivers results within 24–48 hours. This rapid, reliable turnaround shortens decision‑making cycles, lowers travel‑related costs, and improves clinical outcomes for chronic disease management, infectious disease control, and early detection of serious conditions. Looking ahead, integrating AI‑driven decision support, continuous data streams from wearable sensors, and a unified digital health ecosystem will further enhance diagnostic accuracy, enable predictive analytics, and expand equitable access across India’s rural populations. Together, these innovations promise a future where high‑quality, timely care is delivered at the patient’s doorstep, redefining the standard of home healthcare.
Why DNA Testing Matters
Genetic testing is now a cornerstone of modern oncology, enabling clinicians to identify inherited (germline) mutations such as BRCA1/2, TP53, and Lynch‑syndrome genes, as well as somatic alterations that guide targeted therapies and immunotherapy. In India, hereditary cancer syndromes remain markedly under‑diagnosed because of limited access to affordable, accredited genetic services, leading many high‑risk families to miss preventive surveillance and risk‑reducing options. Agam Diagnostics addresses this gap by offering comprehensive medical‑genetics testing—including multigene panels for hereditary risk and tumor DNA profiling—through free home collection, rapid 7‑10‑day turnaround, and full NABL and ICMR accreditation, ensuring high‑quality, reliable results for clinicians and patients alike. These services empower patients to receive surveillance schedules, prophylactic interventions, and eligibility for targeted drugs, ultimately improving outcomes.
Understanding Germline vs. Somatic DNA Testing
Germline (inherited) genetic testing examines DNA from blood or saliva to identify pathogenic variants such as BRCA1/2, TP53, or Lynch‑syndrome genes that raise a person’s lifetime cancer risk. Results guide personalized surveillance, risk‑reducing surgery, and cascade testing of relatives. Somatic testing analyzes DNA extracted from a cancer biopsy or liquid biopsy to detect acquired mutations—e.g., EGFR, KRAS, BRAF, HER2—that drive tumor growth and predict response to targeted therapies or immunotherapies. Clinically, germline testing informs preventive strategies and may uncover eligibility for drugs like PARP inhibitors, while somatic testing directs treatment choices for the current disease, such as EGFR inhibitors in lung cancer or HER2‑targeted therapy in breast cancer. The two approaches are complementary: a somatic panel can reveal a germline‑relevant mutation, prompting germline confirmation, and germline results can affect therapy decisions (e.g., avoiding radiation in TP53 carriers). Together they enable a precision‑medicine workflow that spans risk assessment, early detection, and individualized treatment.
The Role of Genetic Counseling

Genetic counseling is a cornerstone of cancer risk management. Pre‑test risk assessment begins with a detailed three‑generation family history, personal cancer diagnoses, and clinical factors (e.g., early‑onset disease, multiple tumor types) to determine eligibility for germline testing and to select the most relevant gene panel. Interpretation of results is performed by certified counselors who classify findings as pathogenic, likely pathogenic, variant of uncertain significance, or negative, and explain how each category influences surveillance, prophylactic surgery, or targeted therapy (e.g., PARP inhibitors for BRCA‑mutated tumors). Psychosocial support addresses anxiety, potential stigma, and ethical concerns, providing coping strategies and clear communication about insurance protection under regulations such as GINA. Finally, family cascade testing is offered to at‑risk relatives, enabling early detection and preventive interventions across the family unit. Accredited laboratories like Agam Diagnostics (NABL, ICMR) ensure high‑quality results, while free home collection and rapid turnaround facilitate timely counseling and clinical decision‑making.
Biomarker Testing and Targeted Therapy

Companion diagnostic tests pair a specific biomarker assay with an FDA‑approved drug to predict therapeutic effectiveness, turning tumor molecular profiling into a decision‑making tool. Common actionable mutations identified by biomarker testing include EGFR, ALK, KRAS, BRAF, HER2, BRCA1/2, and MSI‑H, each linked to a targeted therapy such as EGFR inhibitors, ALK inhibitors, PARP inhibitors, or HER2‑directed antibodies. Detecting these alterations guides clinicians in selecting the most appropriate regimen—often sparing patients from ineffective chemotherapy and reducing toxicity. Moreover, biomarker results determine eligibility for clinical trials, including basket or umbrella studies, where patients receive novel agents matched to their tumor’s genetic makeup. By integrating companion diagnostics, actionable mutation panels, and trial enrollment criteria, biomarker testing enables truly personalized oncology care.
Personalized Surveillance and Risk‑Reduction Strategies

Individuals who test positive for hereditary cancer mutations benefit from tailored surveillance, prophylactic interventions, chemoprevention, and cascade testing of relatives. Screening schedules are intensified: BRCA1/2 carriers begin annual breast MRI (and mammography after age 30) and ovarian ultrasound or CA‑125 monitoring starting at 35‑40 years, while Lynch‑syndrome carriers undergo colonoscopy every 1–2 years beginning at 20‑25 years and consider endometrial sampling. Prophylactic surgeries offer risk‑reducing options—bilateral mastectomy and salpingo‑oophorectomy dramatically lower breast and ovarian cancer incidence in BRCA mutation carriers, and total colectomy reduces colorectal cancer in high‑penetrance MMR gene families. Chemoprevention, such as tamoxifen for BRCA‑positive women or aspirin for Lynch‑syndrome patients, further attenuates risk. Finally, cascade testing of first‑degree relatives identifies additional mutation carriers, enabling early entry into these surveillance and preventive programs and extending the benefit of the index patient’s genetic insight to the entire family.
Agam Diagnostics: Delivering Quality Genetic Services

Agam Diagnostics, a fully automated pathology laboratory in Madurai, Tamil Nadu, India, combines rigorous accreditation with patient‑centric logistics to provide reliable genetic testing for cancer risk and treatment. The laboratory is accredited by the National Accreditation Board for Testing and Calibration Laboratories (NABL) and the Indian Council of Medical Research (ICMR), guaranteeing compliance with international quality standards, reproducible results, and secure data handling. To remove geographic barriers, Agam offers free home collection of blood or saliva samples, allowing patients in remote or underserved areas to access testing without traveling to a clinic. Once the specimen arrives, next‑generation sequencing (NGS) panels are run on a high‑throughput, fully automated platform, delivering comprehensive analysis of 40‑plus cancer‑related genes within 7‑10 days—a rapid turnaround that supports timely clinical decision‑making. Throughout the process, certified genetic counselors interpret results, explain their clinical significance, and guide patients and their families on surveillance, risk‑reduction, and therapeutic options, ensuring that each test translates into actionable, personalized care.
Overcoming Barriers to Genetic Testing in India

Awareness and education remain the first hurdle; many Indian patients are unaware that 5‑10 % of cancers are hereditary and that testing can guide both prevention and therapy. Community outreach and training of primary‑care physicians can improve referral rates, especially when family‑history questionnaires are used to identify high‑risk individuals. Cultural stigma surrounding cancer and genetic disease often discourages discussion of hereditary risk. Certified genetic counselors can provide culturally sensitive counseling that addresses fears of discrimination and promotes informed decision‑making. Insurance coverage varies widely, but guidelines from NCCN and ASCO recommend reimbursement for germline panels when clinical criteria are met; clear communication between oncologists, counselors, and insurers can increase approval rates. Finally, infrastructure and access are expanding through initiatives such as Agam Diagnostics, which offers free home collection, rapid 7‑10‑day turnaround, and NABL/ICMR‑accredited testing, thereby reaching rural populations and reducing logistical barriers.
Future Directions: Liquid Biopsy, AI, and Clinical Trials

Liquid biopsy offers a minimally invasive way to capture circulating tumor DNA (ctDNA), enabling real‑time monitoring of tumor evolution, early detection of resistance mutations, and access to profiling when tissue is scarce. Coupled with high‑throughput next‑generation sequencing, it can reveal actionable alterations such as EGFR T790M or BRCA reversion mutations, guiding timely therapy switches. Artificial intelligence accelerates the interpretation of these complex genomic datasets, flagging clinically relevant variants, predicting tumor mutational burden, and integrating multi‑omics signals to prioritize treatment options. AI‑driven platforms also streamline patient matching for basket trials, where enrollment is based on a shared biomarker—regardless of tumor site—expanding access to targeted agents for rare alterations like NTRK fusions or high microsatellite instability. Emerging biomarkers, including tumor‑derived exosomal RNA, epigenetic signatures, and proteomic panels, promise to refine risk stratification and therapeutic selection, further cementing liquid biopsy, AI, and adaptive trial designs as pillars of next‑generation precision oncology.
Putting DNA Testing into Practice
Genetic testing delivers concrete benefits: it pinpoints inherited cancer‑predisposition mutations (e.g., BRCA1/2, Lynch genes), enables earlier, intensified surveillance, informs risk‑reducing surgeries, and guides tumor‑targeted therapies such as PARP or EGFR inhibitors. Patients with strong family histories, early‑onset disease, or rare syndromes should discuss testing with a certified genetic counselor, and providers must integrate testing into routine oncology workflows. Timely ordering of germline and somatic panels—ideally through accredited facilities—guarantees reliable, high‑quality results that meet international standards (NABL, ICMR). Accredited labs such as Agam Diagnostics offer free home collection and rapid turnaround, removing barriers and ensuring that accurate DNA insights translate into personalized prevention and treatment plans.
A New Era for Blood‑Disorder Care
The rapid expansion of cellular therapies, from CAR‑T cells (liso‑cel, ide‑cel) to bispecific antibodies, is reshaping treatment algorithms for B‑cell lymphomas, follicular lymphoma, and CNS‑involved myeloma, delivering response rates up 85% and durable remissions. Simultaneously, automation and AI‑driven hematology analyzers—integrating laser‑based flow cytometry, digital morphology, and machine‑learning flagging—are cutting turnaround times to minutes, enabling real‑time MRD monitoring and early detection of abnormal cells. Precision medicine is now anchored by comprehensive genomic profiling via next‑generation sequencing panels that uncover actionable mutations (e.g., BTK, BCL‑2, JAK‑STAT) and guide targeted regimens such as acalabrutinib + venetoclax or pelabresib + ruxolitinib. Together, these advances create a seamless pipeline from rapid, AI‑enhanced diagnostics to personalized, cellular‑based therapeutics, accelerating outcomes for patients with hematologic disorders.
CAR‑T Cells and Bispecific Antibodies Move to Earlier Lines

In 2024, CAR‑T and bispecific modalities are shifting from salvage to frontline use. Real‑world CIBMTR data (n = 156) showed lisocabtagene maraleucel (liso‑cel; Breyanzi) achieved a 6‑month PFS of 61 % and OS of 87 % as second‑line therapy for relapsed/refractory large B‑cell lymphoma, while the ELARA phase‑2 update reported tisagenlecleucel with a 4‑year OS of 79.3 % and a 69.1 % CR rate in follicular lymphoma. Dual‑target CD20/CD19 antibodies further lowered progression and death risk in follicular lymphoma (ASH LBA‑1). Bispecific T‑cell engagers are also delivering striking results: blinatumomab raised three‑year disease‑free survival to 97 % in pediatric B‑ALL, and epcoritamab produced a 61 % overall response rate in relapsed/refractory CLL after BTK and BCL‑2 inhibitor failure. These outcomes underscore the expanding role of immunotherapy earlier in treatment algorithms.
AI‑Driven Automation Accelerates Hematology Testing

Mindray’s newest hematology analyzers marry laser‑based flow cytometry, impedance and fluorescence to generate detailed cell morphology, Immature Platelet Fraction (IPF) and optical platelet counts in seconds. The integrated AI algorithms automatically flag abnormal cells, calculate derived parameters such as Reticulocyte Hemoglobin (RET‑He) and inflammation ratios, and dramatically cut the need for manual slide review. In parallel, digital morphology platforms like Mindray’s MC‑80 scan peripheral‑smear slides, apply deep‑learning models to detect blasts, atypical lymphocytes and other subtle changes, and feed the results directly into the laboratory information system. Agam Diagnostics in Madurai has built a fully automated, 24‑hour workflow around these technologies, offering free home collection, NABL and ICMR accreditation, and a turnaround‑ under 24 hours for complete blood counts and specialized hematology panels. Together, AI‑driven analysis and end‑to‑end automation are reshaping laboratory hematology by delivering faster, more accurate diagnoses while freeing technologists for higher‑value tasks.
Next‑Generation Sequencing Powers Precision Medicine

Comprehensive next‑generation sequencing (NGS) panels have become a cornerstone of modern hematology, especially for AML, MDS, and CLL. By simultaneously detecting mutations in genes such as FLT3, NPM1, DNMT3A, IDH1/2, and a broad array of epigenetic regulators, NGS enables clinicians to match patients with targeted agents and determine eligibility for novel therapies. In acute myeloid leukemia, the molecular risk signature (mPRS) stratifies individuals receiving hypomethylating agents plus venetoclax into three outcome groups, guiding intensity of treatment and informing prognosis. In mantle‑cell lymphoma, NGS‑enhanced minimal residual disease (MRD) assessment allows treatment de‑escalation, permitting omission of high‑dose chemotherapy and autologous stem‑cell transplant without compromising long‑term survival. These advances illustrate how NGS directly influences treatment selection, risk stratification, and therapeutic de‑intensification across hematologic malignancies.
Oral Targeted Agents and BTK Degraders Redefine CLL Therapy

Oral agents are reshaping chronic lymphocytic leukemia (CLL) management. Five‑year SEQUOIA data demonstrate that zanubrutinib (Brukinsa) delivers superior progression‑free survival and a more favorable safety profile than bendamustine‑rituximab in treatment‑naïve CLL/SLL, establishing it as a new standard of care. The AMPLIFY phase‑3 trial showed that a fixed‑duration, all‑oral regimen of acalabrutinib plus venetoclax (doublet) achieved an 83.1% 36‑month PFS, markedly outpacing standard chemo‑immunotherapy. Emerging BTK degraders, exemplified by BGB‑16673, have reported an overall response rate of ~78% in relapsed/refractory CLL, offering a strategy to overcome resistance to conventional BTK inhibitors. Together, these oral therapies and novel degraders are redefining the CLL treatment paradigm by providing highly effective, chemotherapy‑free options with durable responses and improved tolerability.
Gene Therapy and CRISPR Move Toward Curative Hemoglobinopathies

Recent advances have turned gene‑editing into a realistic cure for hemoglobin disorders. Exagamglogene autotemcel (Exa‑cel) became the first FDA‑approved ex vivo CRISPR‑Cas9 therapy for sickle cell disease, editing the BCL11A enhancer in autologous hematopoietic stem cells to reactivate fetal hemoglobin and achieve durable transfusion‑independence. Lovotibeglogene autotemcel, a lentiviral vector that adds a functional β‑globin gene (HbAT87Q), received approval for sickle cell disease and expands the curative toolkit for both sickle cell and β‑thalassemia. Parallel pre‑clinical work is testing in‑vivo CRISPR delivery—using adeno‑associated viral capsids or lipid‑nanoparticle carriers—to edit hematopoietic stem cells directly in patients, which could eliminate the need for cell collection and ex‑vivo manipulation. These milestones collectively illustrate that precise genome‑editing is rapidly moving from experimental to therapeutic reality for hemoglobinopathies.
Lifestyle, Supportive Care, and MRD‑Based Strategies

Non‑pharmacologic factors are emerging as powerful modifiers of hematologic outcomes. Current pre‑clinical data (ASH abstract 259) show that higher dietary fiber intake after allogeneic stem‑cell transplantation attenuates gastrointestinal graft‑versus‑host disease and translates into better overall survival, likely through modulation of gut microbiota and inflammatory pathways. Conversely, smoking exerts a dose‑responsive increase in somatic mutational burden, accelerating the evolution of myelodysplastic syndrome to acute myeloid leukemia (ASH abstract 4597, underscoring the need for aggressive smoking cessation programs in at‑risk populations. In mantle‑cell lymphoma, minimal residual disease (MRD) monitoring now guides treatment de‑escalation: patients with MRD‑negative status can safely skip high‑dose chemotherapy and autologous stem‑cell transplant without compromising long‑term survival (ASH abstract LBA‑6. Together, these findings highlight how lifestyle interventions and MRD‑driven decision‑making can personalize care, reduce treatment toxicity, and improve survival across diverse hematologic diseases.
Looking Ahead: Integrating Innovation with Accessible Care
Automated laboratories such as Agam Diagnostics are already delivering cutting‑edge haematology testing—AI‑driven differential counts, rapid CBCs, and advanced flow‑cytometry panels—within 24 hours while offering free home collection and NABL/ICMR‑accredited quality. Scaling these capabilities nationwide will help India’s 1.5 million‑strong Madurai region and other underserved areas access molecular panels, MRD monitoring, and NGS‑based mutation profiling essential for personalized therapy. Equitable distribution of novel agents—CAR‑T cells, bispecific antibodies, oral BTK degraders, and next‑generation gene‑editing treatments—requires public‑funded reimbursement, regional trial sites, and tele‑pathology networks. Looking forward, AI‑assisted diagnostics, CRISPR‑based curative approaches, and fixed‑duration oral regimens (e.g., acalabrutinib + venetoclax) will further bridge the gap between breakthrough science and everyday patient care across India.


























