Why Microbiology Matters in Emerging Disease Surveillance
Surveillance is the continuous, systematic collection, analysis and interpretation of health data to guide public‑health planning, implementation and evaluation. Clinical microbiology laboratories have been the first point of detection for infectious threats since the advent of culture and microscopy, evolving from simple Gram stains to sophisticated molecular platforms that can sequence entire pathogen genomes in hours. Emerging threats—novel viruses such as SARS‑CoV‑2, influenza A variants, bioterrorism agents, and zoonotic spillovers like Nipah or Streptococcus suis—underscore the need for rapid, accurate laboratory data. Modern labs, exemplified by fully automated facilities such as Agam Diagnostics in Madurai, provide same‑day PCR, whole‑genome sequencing, and automated alert systems that feed into regional, national and international surveillance networks, enabling early intervention and containment.
Surveillance Foundations and the Role of Microbiology Laboratories

Surveillance is the continuous, systematic collection, analysis, and interpretation of health data that underpins public‑health planning, implementation, and evaluation.
Clinical microbiology laboratories serve as the first detection point for emerging threats and feed data into three surveillance models. Passive surveillance relies on routine, often paper‑based, analysis of all microbiological results, generating alerts after the fact. Passive surveillance relies on routine analysis of all microbiological data and is often paper‑based. Active surveillance applies predefined thresholds and engages trained microbiologists and epidemiologists to seek out abnormal patterns, enabling faster outbreak detection. Active surveillance applies predefined thresholds and engages trained microbiologists and epidemiologists. Virtual surveillance combines active data collection with advanced computing, mathematical models, and real‑time analytics to flag atypical pathogen trends instantly. Virtual surveillance combines active surveillance with advanced computing technologies and mathematical models to provide real‑time detection of abnormal pathogen patterns. Automated laboratory information systems in microbiology labs generate electronic alerts, stream‑ data to regional, national, and international networks such as WHO GLASS, WHONET, and national Integrated Disease Surveillance Programs. Automated data management systems in microbiology labs can generate alerts for unusual results, detect trends, and integrate data into regional, national, and international surveillance networks. By integrating specimen management, rapid molecular diagnostics, and robust data pipelines, microbiology laboratories enhance temporal and spatial disease mapping, support early intervention, and improve coordinated public‑health responses worldwide.
Emerging Pathogens and the Need for Rapid Detection

Rapid detection of emerging pathogens such as SARS, novel influenza A variants, bioterrorism agents, and zoonotic spillovers is now a public‑health imperative. Delayed recognition allows unchecked transmission, inflating outbreak size and mortality; the 2002‑2003 SARS epidemic and the 2001 anthrax mail attacks both demonstrated how weeks of diagnostic lag amplified case numbers and panic. Recent laboratory data illustrate the stakes: SARS‑CoV‑2 has been isolated from white‑tailed deer in Ohio, indicating wildlife reservoirs that can reseed human infections if not identified early. Likewise, the emergence of OXA‑48‑producing Escherichia coli in food‑premises highlights how antibiotic‑resistant bacteria can spread silently through the food chain, evading routine culture‑based surveillance. Real‑time PCR, whole‑genome sequencing, and automated data‑alert systems—implemented in accredited facilities such as Agam Diagnostics—provide same‑day pathogen identification, resistance‑gene profiling, and rapid reporting to national and international networks. These technologies shrink turnaround times from days to hours, enabling timely isolation, targeted therapy, and coordinated public‑health responses that curb the magnitude of emerging infectious disease threats.
Molecular Diagnostics: PCR, Real‑time PCR, and Next‑Generation Sequencing
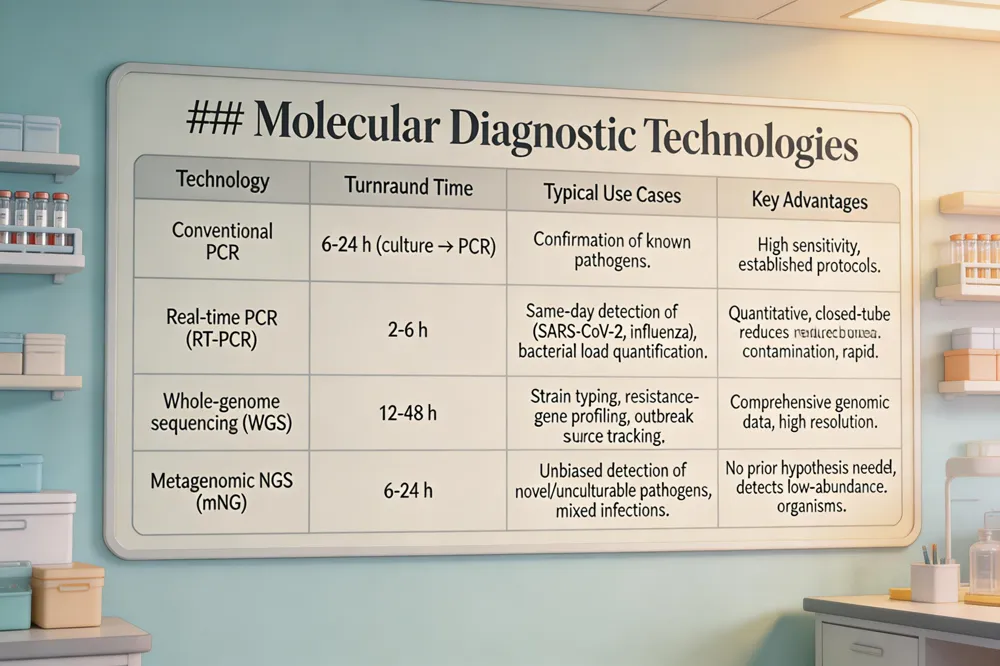

Real‑time PCR (RT‑PCR) amplifies nucleic acids while simultaneously measuring fluorescence, delivering quantitative results in 2‑6 hours. Its closed‑tube format minimizes contamination and, combined with automated extraction, shortens turnaround from days (culture) to hours, enabling same‑day detection of emerging viruses such as SARS‑CoV‑2, influenza A variants, and zoonotic agents. Whole‑genome sequencing (WGS) and metagenomic next‑generation sequencing (mNGS) go further by reading the entire genetic material of a specimen without prior culturing. These platforms identify novel or unculturable pathogens, map mutations that affect virulence, and reveal transmission pathways in real time. In antimicrobial‑resistance surveillance, WGS pinpoints resistance genes (e.g., OXA‑48, mcr‑1) and plasmid backgrounds, while mNGS can detect low‑abundance resistant strains in mixed infections. Together, RT‑PCR provides rapid, targeted alerts for known threats, whereas WGS/mNGS supplies comprehensive genomic data for strain tracking, outbreak source attribution, and informed public‑health interventions.
Automation and Core Laboratory Models for Speed and Cost‑Effectiveness

Horizontal core laboratory structures bring microbiology, immunology and biochemistry under a single, unified workflow, eliminating duplicated steps and streamlining staffing. By consolidating these disciplines, laboratories can share high‑throughput analyzers, automated incubators and common data‑management platforms, which reduces capital outlay and improves turnaround time for all test types. Robotic specimen handling—cover bar‑code‑driven sorting, automated aliquoting and sealed biosafety cabinets—paired with an integrated Laboratory Information System (LIS) enables real‑time data capture, trend detection and automatic alert generation for unusual results. These alerts feed directly into regional and national surveillance networks, shortening reporting from days to hours. In high‑volume settings such as Agam Diagnostics in Madurai, the model yields measurable cost savings through lower consumable waste, fewer repeat tests and optimized personnel allocation, while delivering same‑day pathogen identification and antimicrobial‑susceptibility results essential for outbreak control.
Specimen Management and Biosafety Best Practices

Effective specimen management begins with proper collection: trained staff must use sterile containers, follow pathogen‑specific protocols, and label each sample with patient identifiers, date, time, and test request. Immediate, temperature‑controlled transport—refrigerated for most bacterial and viral specimens, frozen for RNA viruses—preserves nucleic‑acid integrity, while prompt storage at the lab prevents degradation. In the microbiology laboratory, Class II biosafety cabinets provide a protected environment for handling potentially infectious material; personnel must wear appropriate personal protective equipment (gloves, lab coats, eye protection, and N95 respirators when aerosol‑generating procedures are performed. Waste is decontaminated by autoclaving or chemical disinfection before disposal. High‑quality specimens are essential: contaminated or poorly preserved samples yield false‑negative results, undermine antimicrobial‑resistance surveillance, and delay outbreak detection. By standardising collection, transport, and biosafety measures, laboratories such as Agam Diagnostics ensure accurate diagnostics and reliable data for public‑health surveillance.
Data Integration, Automated Alerts, and Virtual Surveillance
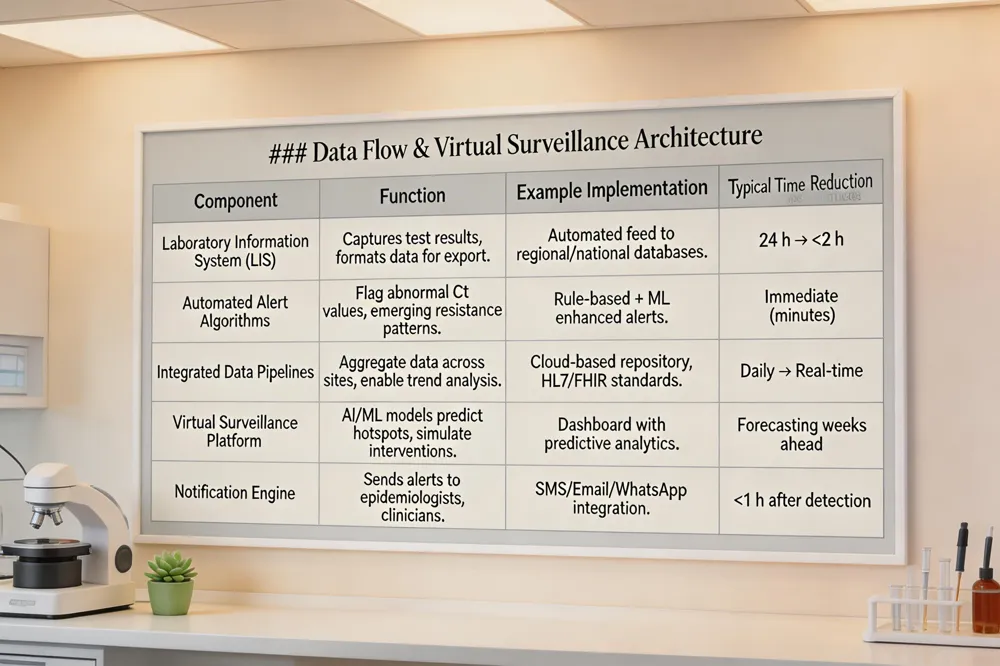

Modern clinical microbiology laboratories, such as Agam Diagnostics in Madurai, now employ robust Laboratory Information Systems (LIS) that automatically feed test results into regional, national, and supranational surveillance networks (e.g., WHO GLASS, WHONET). These electronic links enable near‑real‑time aggregation of pathogen data, allowing automated alert algorithms to flag unusual results—such as unexpected viral PCR Ct values or emergence of carbapenem‑resistant bacterial isolates—within hours of receipt. Integrated data pipelines support trend analysis across time and space, facilitating early outbreak detection and targeted public‑health interventions. Building on this foundation, virtual surveillance platforms incorporate mathematical modeling and artificial‑intelligence tools to predict abnormal pathogen patterns, assess transmission dynamics, and simulate the impact of control measures. AI‑driven dashboards can prioritize alerts, suggest laboratory actions, and automatically notify epidemiologists and clinicians, thereby shortening reporting time from a median of five days to as little as one day. Together, LIS connectivity, automated alerts, and AI‑backed virtual surveillance create a rapid, data‑centric defense against emerging infectious disease threats.
Agam Diagnostics: Infrastructure, Accreditation, and Service Reach

Agam Diagnostics is a fully automated pathology laboratory situated in Madurai, Tamil Nadu, India. It operates a high‑throughput, integrated platform that combines microbiology, molecular biology, haematology, clinical biochemistry, immunology and medical genetics under one roof, enabling same‑day or next‑day reporting for most assays. The laboratory holds NABL accreditation (ISO 15189:2022) and is also accredited by the Indian Council of Medical Research (ICMR), ensuring compliance with stringent quality‑assurance programmes, regular participation in external quality‑assessment schemes, and a documented 99.9 % accuracy rate for test results. To maximise accessibility, Agam Diagnostics offers free home‑sample collection across Tamil Nadu and a network of partner laboratories in seven additional locations, ensuring timely specimen transport and rapid turnaround even for patients in remote areas. This combination of automation, accreditation, and outreach positions the lab as a critical node for early detection of emerging infectious diseases and for feeding reliable data into national surveillance systems.
Impact on Public Health: Early Intervention, Outbreak Control, and Antimicrobial Resistance Monitoring

Temporal and spatial analysis of disease patterns becomes possible when clinical microbiology laboratories feed high‑quality, time‑stamped data into national surveillance networks. Automated laboratory information systems generate alerts for unusual pathogen trends, allowing health authorities to map hotspots and predict spread across districts. Rapid antimicrobial susceptibility testing (AST) shortens the interval from specimen receipt to therapeutic decision: hours rather than days, enabling clinicians to prescribe targeted agents and curb the use of broad‑spectrum antibiotics that drive resistance. In India, laboratories such as Agam Diagnostics, which are NABL and ICMR accredited, integrate their AST results with the Integrated Disease Surveillance Programme (IDSP) and the WHO Global Antimicrobial Resistance Surveillance System (GLASS). This linkage provides real‑time resistance profiles at the regional and national levels, supporting evidence‑based treatment guidelines, informing infection‑control policies, and enhancing early outbreak response. The combined effect is a more agile public‑health system that can intervene promptly, limit transmission, and monitor antimicrobial resistance trends effectively.

Unbiased pathogen detection through shotgun metagenomics will soon become routine in microbiology labs such as Agam Diagnostics, allowing clinicians to identify known, novel, or unculturable agents directly from clinical specimens without prior hypotheses. Coupled with rapid next‑generation sequencing platforms, this approach can pinpoint viral variants, antimicrobial‑resistance genes, and zoonotic spill‑over events within hours, feeding real‑time data into regional and national surveillance networks. Machine‑learning algorithms will ingest these high‑dimensional datasets, together with electronic laboratory reporting and epidemiological indicators, to predict outbreak hotspots, forecast resistance trends, and generate automated alerts for unusual pathogen signatures. The One Health model further expands the data stream by integrating human, animal, and environmental microbiology results—such as wildlife sampling for SARS‑CoV‑2 variants or livestock testing for emerging bacterial strains—into a unified analytical framework. Together, metagenomics, AI, and One Health surveillance promise faster, more precise public‑health responses to emerging infectious diseases.
Putting It All Together – The Critical Role of Microbiology in a Changing Infectious‑Disease Landscape
Clinical microbiology laboratories provide the first line of detection for emerging threats, turning specimen collection, culture, microscopy, antigen assays, and molecular tests into actionable data. Rapid diagnostics—real‑time PCR, next‑generation sequencing, MALDI‑TOF—identify pathogens within hours, allowing clinicians to start appropriate therapy and public‑health agencies to trigger outbreak alerts. Agam Diagnostics exemplifies this modern model: a NABL‑ and ICMR‑accredited, fully automated facility in Madurai that offers bacterial culture, viral PCR panels, antimicrobial‑susceptibility testing, and free home sample collection. Its integrated laboratory information system generates electronic alerts and feeds results into regional surveillance networks, shortening reporting times from days to a single day. To sustain these gains, continued investment is needed in cutting‑edge platforms, regular staff training, and robust data‑sharing frameworks that link microbiology labs with epidemiologists and national surveillance programs. Such support will keep the laboratory’s early‑warning capability sharp as pathogens evolve and spread.